DNA Replication: How DNA Copies Itself
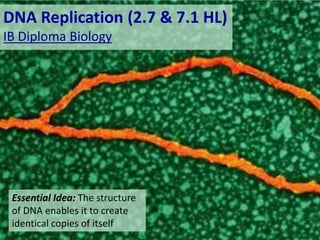
- 1. DNA Replication (2.7 & 7.1 HL) IB Diploma Biology Essential Idea: The structure of DNA enables it to create identical copies of itself
- 2. 2.7.1 The replication of DNA is semi-conservative and depends on complimentary base-pairing.
- 3. Semi-Conservative? 2.7.1 The replication of DNA is semi-conservative and depends on complimentary base-pairing. DNA replication is…
- 4. Semi-conservative? 2.7.1 The replication of DNA is semi-conservative and depends on complimentary base-pairing.
- 5. 2.7.1 The replication of DNA is semi-conservative and depends on complimentary base-pairing.
- 6. Each time DNA is copied, the new double stranded molecule consists of one old template strand plus a new complementary strand made from previously free bases 2.7.1 The replication of DNA is semi-conservative and depends on complimentary base-pairing.
- 7. A = T and C = G, they are complementary The way the molecules fit together makes it very unlikely that they will bond with the wrong partner. So the genetic code is faithfully copied during replication When things do go wrong, we have a mutation! 2.7.1 The replication of DNA is semi-conservative and depends on complimentary base-pairing.
- 8. 2.7.12 Analysis of Meselson and Stahl’s results to obtain support for the theory of semi-conservative replication of DNA / 7.1.1 DNA structure suggested a mechanism for DNA replication. • Watson & Crick’s 1953 paper on DNA structure ended by suggesting a semi-conservative model for DNA replication… • Matthew Meselson and Franklin Stahl designed what is often called the “most beautiful experiment in biology” to prove their hypothesis about DNA replication http://www.learnerstv.com/animation/biology/con20ani.swf
- 9. 2.7.12 Analysis of Meselson and Stahl’s results to obtain support for the theory of semi-conservative replication of DNA. http://highered.mheducation.com/olcweb/cgi/pluginpop.cgi?it=swf::535 ::535::/sites/dl/free/0072437316/120076/bio22.swf::Meselson%20and %20Stahl%20Experiment • E. coli bacteria were grown in two different isotopes of Nitrogen: • Bacteria grown in N-14 isotope had ‘lighter’ DNA • Bacteria grown in N-15 isotope had ‘heavier’ DNA • N-15 bacteria were transferred into N- 14 solution and samples were collected every 20 minutes (time of E. coli replication cycle) • Samples were centrifuged to separate DNA by weight (see right)
- 10. Helicase • The ‘-ase’ ending indicates it is an enzyme • This family of proteins varies, but are often formed from multiple polypeptides and doughnut in shape 2.7.2 Helicase unwinds the double helix and separates the two strands by breaking hydrogen bonds.
- 11. • Unwinds the DNA Helix • Separates the two polynucleotide strands by breaking the hydrogen bonds between complementary base pairs • ATP is needed by helicase to both move along the DNA molecule and to break the hydrogen bonds • The two separated strands become parent / template strands for the replication process 2.7.2 Helicase unwinds the double helix and separates the two strands by breaking hydrogen bonds.
- 12. DNA Polymerase • The ‘-ase’ ending indicates it is an enzyme • This protein family consists of multiple polypeptides • The polymerization reaction is a condensation reaction 2.7.3 DNA polymerase links nucleotides together to form a new strand, using the pre-existing strand as a template.
- 13. • Free nucleotides are nucleoside triphosphates • The extra phosphate groups carry energy which is used for formation of covalent bonds • DNA polymerase always moves in a 5’ to 3’ direction • DNA polymerase catalyzes the covalent Phosphodiester bonds between sugars and phosphates • DNA Polymerase proofreads the complementary base pairing so mistakes are very rare, occurring approx. once in every billion bases 2.7.3 DNA polymerase links nucleotides together to form a new strand, using the pre-existing strand as a template.
- 14. 2.7.3 DNA polymerase links nucleotides together to form a new strand, using the pre-existing strand as a template.
- 15. The Polymerase Chain Reaction (PCR) Synthetic method of amplifying specific sequences of DNA. Useful when only a small amount of DNA is available for testing e.g. crime scene samples of blood, semen, hair, etc. http://highered.mcgraw- hill.com/olc/dl/120078/micro15.swf 2.7.9 Use of Taq DNA polymerase to produce multiple copies of DNA rapidly by the polymerase chain reaction (PCR). • Processes artificially recreates DNA replication • Taq DNA Polymerase is used for PCR • Comes from heat-resistant bacterium, Thermus aquaticus, that lives in hot springs… • Can resist denaturation at high temperatures required to separate DNA strands in PCR • Copies up to 1000 nucleotides / minute
- 16. The PCR Process: PCR occurs in a thermal cycler and involves 3 steps: 1. Denaturation: DNA sample is heated to 95⁰C to break hydrogen bonds and separate it into two strands 2. Annealing: DNA sample is cooled to 54 ⁰C, allowing primers attach to opposite ends of the target sequence 3. Elongation: A heat-tolerant DNA polymerase (Taq) copies the strands • One cycle of PCR yields two identical copies of the DNA sequence • A standard reaction of 30 cycles would yield 1,073,741,826 copies of DNA (230) 2.7.9 Use of Taq DNA polymerase to produce multiple copies of DNA rapidly by the polymerase chain reaction (PCR).
- 17. 2.7.9 Use of Taq DNA polymerase to produce multiple copies of DNA rapidly by the polymerase chain reaction (PCR).
- 18. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer. DNA Polymerase can only add nucleotides to the 3’ end…
- 19. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer.
- 20. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer.
- 21. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer.
- 22. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer. http://highered.mheducation.com/olcweb/cgi/pluginpop.cgi?it=swf::535::535::/sites/dl/free/0072437316 /120076/bio23.swf::How%20Nucleotides%20are%20Added%20in%20DNA%20Replication
- 23. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer.
- 24. 7.1.3 DNA replication is continuous on the leading strand and discontinuous on the lagging strand / 7.1.5 DNA polymerases can only add nucleotides to the 3’ end of a primer.
- 25. 7.1.4 DNA replication is carried out by a complex series of enzymes. DNA Polymerase III DNA Gyrase (Topoisomerase) stabilizes the helix as it is unwound
- 26. 7.1.4 DNA replication is carried out by a complex series of enzymes.
- 27. 7.1.4 DNA replication is carried out by a complex series of enzymes.
- 28. 7.1.4 DNA replication is carried out by a complex series of enzymes.
- 29. Enzyme Function DNA Gyrase (Topoisomerase) Stabilizes DNA helix as it is unwound by Helicase Single Stranded Binding Proteins Hold unzipped, single-stranded sections of DNA apart during replication Helicase Unwinds DNA helix and unzips strands by breaking Hydrogen bonds DNA Polymerase III Adds new DNA nucleotides in the 5’ 3’ direction DNA Primase Adds primers of RNA nucleotides to the lagging strand as starting points for replication DNA Polymerase I Replaces RNA primers with DNA nucleotides DNA Ligase Joins Okazaki fragments on lagging stand 7.1.4 DNA replication is carried out by a complex series of enzymes.
- 30. 7.1.4 DNA replication is carried out by a complex series of enzymes. http://www.stolaf.edu/people/giannini/flas hanimat/molgenetics/dna-rna2.swf http://sites.fas.harvard.edu/~biotext/ animations/replication1.swf
- 31. 7.1.9 Use of nucleotides containing dideoxyribonucleic acid to stop DNA replication in preparation of samples for base sequencing. DNA Sequencing: • Unknown DNA sequence mixed with DNA nucleotides and enzymes needed for replication • Dideoxyribnonucleotides with different fluorescent markers are added • These modified nucleotides stop replication at the point they are added to the DNA since new nucleotides cannot be attached to their 3’ end • Fragments are separated by size and analyzed by fluorescence
- 32. 7.1.9 Use of nucleotides containing dideoxyribonucleic acid to stop DNA replication in preparation of samples for base sequencing.
- 33. Bibliography / Acknowledgments Jason de Nys Chris Paine
